Visuals
UMAP Embedding Projections
Each plot shows 64-dimensional embeddings reduced to 2D via UMAP. Points are colored by ground-truth class labels. Tight, well-separated clusters indicate that the embedding captures the graph's community structure.
Cora

Cleora 86.1%
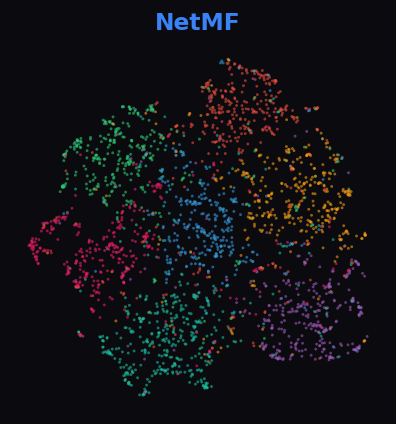
NetMF 83.9%

DeepWalk 83.5%

HOPE 82.1%

ProNE 17.9%

RandNE 24.7%
Cleora shows the tightest, most distinct clusters — 7 classes cleanly separated. NetMF and DeepWalk also show structure but with more overlap. ProNE and RandNE produce noisy, unstructured projections.
CiteSeer
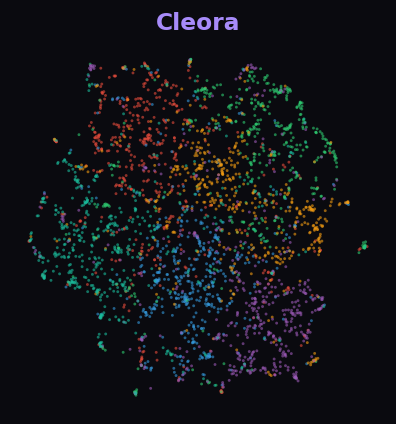
Cleora 82.4%
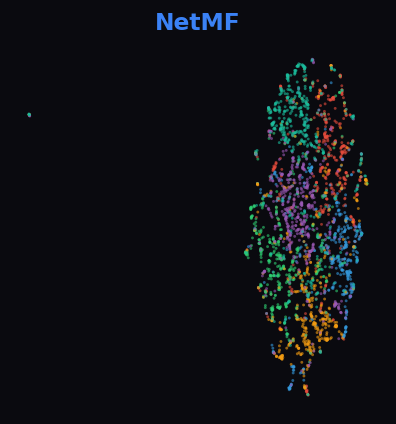
NetMF 81.0%

HOPE 72.4%

ProNE 21.1%

RandNE 20.2%
Cleora separates 6 subject areas into clear, compact groups. NetMF is close behind but with more inter-class mixing. HOPE captures some structure. ProNE and RandNE show little meaningful separation.

Cleora 99.0%

NetMF 95.7%

ProNE 68.5%
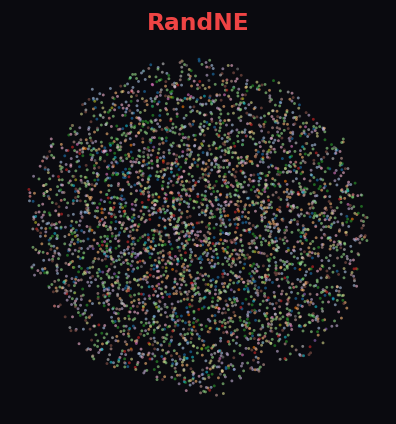

RandNE 63.2%
Facebook ego-network communities detected via Louvain. Cleora and NetMF both capture the community structure well, with Cleora achieving the tightest separation. ProNE and RandNE show less organized projections.
PubMed
Only 3 algorithms complete on PubMed — HOPE, NetMF, GraRep, DeepWalk, and Node2Vec fail (timeout or OOM).
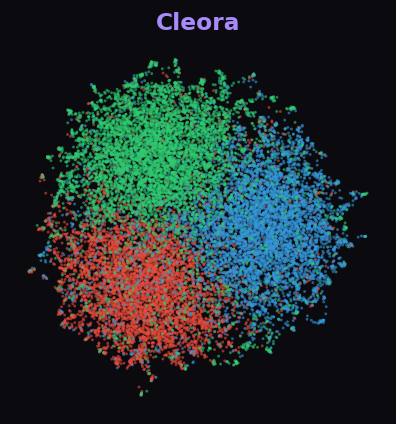
Cleora 87.9%
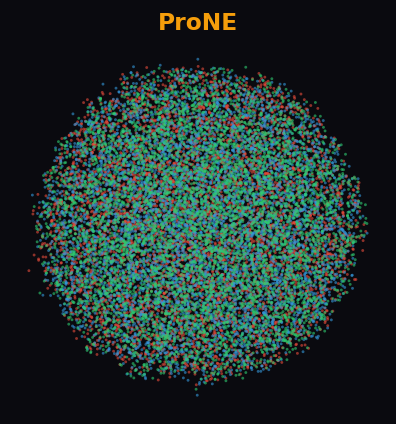

ProNE 33.9%
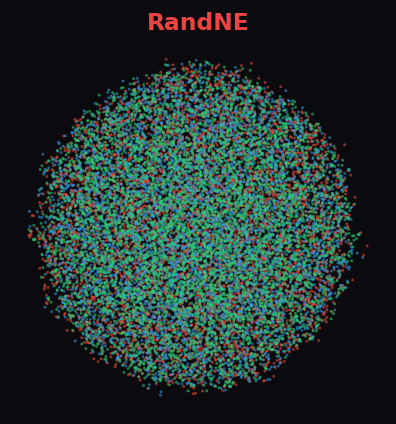

RandNE 35.1%
At 19.7K nodes, most algorithms crash. Cleora cleanly separates all 3 diabetes paper categories with distinct, well-formed clusters. ProNE and RandNE produce uniform noise — the difference is dramatic.
PPI
Only 3 algorithms complete on PPI — HOPE, NetMF, GraRep, DeepWalk, and Node2Vec fail (timeout or OOM).

Cleora 100%

ProNE 2.3%

RandNE 7.3%
PPI has 50 protein function classes. Cleora achieves perfect classification accuracy and its UMAP shows rich, multi-cluster structure. ProNE and RandNE produce near-random noise with no discernible class separation.
Methodology
- All embeddings use 64 dimensions, reduced to 2D via UMAP (
n_neighbors=15,min_dist=0.1,random_state=42) - Cleora uses
num_iterations=40,propagation='left',normalization='l2',whiten=True - Points are colored by ground-truth labels (Cora/CiteSeer/PubMed/PPI) or Louvain communities (Facebook)
- Accuracy numbers from the full benchmark suite (separate 256-dim runs, 3-run average)